HowTo¶
This section describes how to run each analysis function independently.
Calculate similarity¶
The example code:
"""
Created on Sat Apr 3 23:48:38 2021
@author: arsi
"""
import pandas as pd
from tscfat.Analysis.calculate_similarity import calculate_similarity
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# convert one column to a numpy array
name = 'positive'
ser = df[name].values.reshape(-1,1)
sim = calculate_similarity(ser)
The similarity matrix:
print(sim)
[[1. 0.8962963 0.8962963 ... 0.78317152 0.75467775 0.81208054]
[0.8962963 1. 0.81208054 ... 0.86120996 0.82687927 0.8962963 ]
[0.8962963 0.81208054 1. ... 0.71810089 0.69407266 0.74233129]
...
[0.78317152 0.86120996 0.71810089 ... 1. 0.95400788 0.95652174]
[0.75467775 0.82687927 0.69407266 ... 0.95400788 1. 0.91435768]
[0.81208054 0.8962963 0.74233129 ... 0.95652174 0.91435768 1. ]]
Calculate Novelty¶
The example code:
"""
Created on Sat Apr 3 23:48:38 2021
@author: arsi
"""
import pandas as pd
from tscfat.Analysis.calculate_similarity import calculate_similarity
from tscfat.Analysis.calculate_novelty import compute_novelty
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# convert one column to a numpy array
name = 'positive'
ser = df[name].values.reshape(-1,1)
# similarity matrix
sim = calculate_similarity(ser)
# novelty score and kernel
nov, kernel = compute_novelty(sim,edge=7)
The novelty score:
print(nov)
[0.20766337 0.14769835 0.10181832 0.06656327 0.04818483 0.02417862
0.01317275 0.01011372 0.00741401 0.00831496 0.00726132 ...
... 0.01400025 0.03109473 0.06095036 0.10315707 0.16478283 0.23854444]
Calculate Stability¶
The example code:
"""
Created on Sat Apr 3 23:48:38 2021
@author: arsi
"""
import pandas as pd
from tscfat.Analysis.calculate_similarity import calculate_similarity
from tscfat.Analysis.calculate_stability import compute_stability
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# convert one column to a numpy array
name = 'positive'
ser = df[name].values.reshape(-1,1)
# similarity matrix
sim = calculate_similarity(ser)
# stability score
stab = compute_stability(sim)
The stability score:
print(stab)
[0. 0. 0. 0.66850829 0.76100629 0.81208054
0.84615385 0.8705036 0.88321168 0.88321168 0.8705036 ...
... 0.93316195 0.93556701 0.93556701 0.93556701 0.92602041
0.90636704 0.87893462 0.8705036 0.84320557 0. 0.
0. ]
Plot Similarity¶
The example code:
"""
Created on Sat Apr 3 23:48:38 2021
@author: arsi
"""
import pandas as pd
from tscfat.Analysis.calculate_similarity import calculate_similarity
from tscfat.Analysis.calculate_novelty import compute_novelty
from tscfat.Analysis.calculate_stability import compute_stability
from tscfat.Analysis.plot_similarity import plot_similarity
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# convert one column to a numpy array
name = 'positive'
ser = df[name].values.reshape(-1,1)
# kernel edge length -> window = 2*edge + 1
edge = 7
# similarity, stability and novelty
sim = calculate_similarity(ser)
stab = compute_stability(sim)
nov, kernel = compute_novelty(sim, edge = edge)
_ = plot_similarity(sim,
nov,
stab,
title = "Positive affect Similarity, Novelty and Stability",
doi = None,
savepath = False,
savename = False,
ylim = (0,0.05),
threshold = 0.9,
axis = None,
kernel = kernel,
test = False
)
The output image:

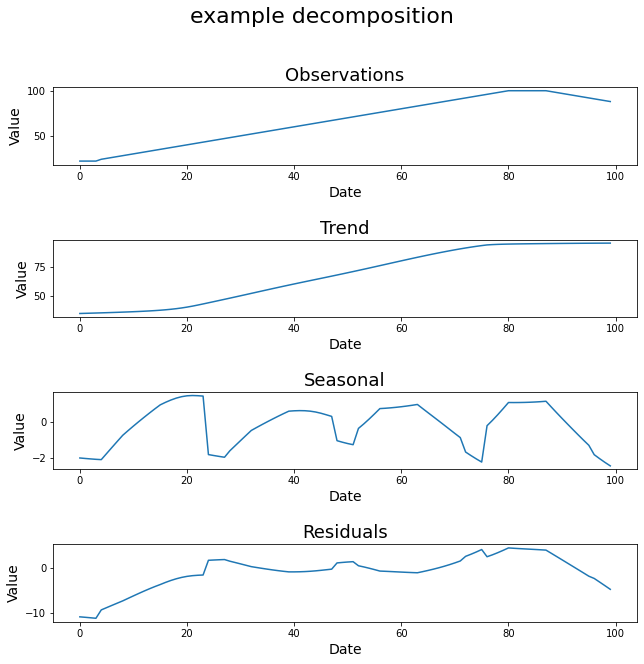
Timeseries Decomposition¶
STL_decomposition function decomposes the given timeseries into trend, seasonal, and residual components.
The example code:
"""
Created on Thu Apr 1 14:41:53 2021
@author: arsii
"""
import pandas as pd
from tscfat.Analysis.decompose_timeseries import STL_decomposition
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsii/tscfat/Data/Test_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# timeseries has to be a numpy array
series = df.level.values
_ = STL_decomposition(series,
title = 'example decomposition',
test = False,
savepath = False,
savename = False,
ylabel = "Value",
xlabel = "Date",
dates = False,
)
The output image:
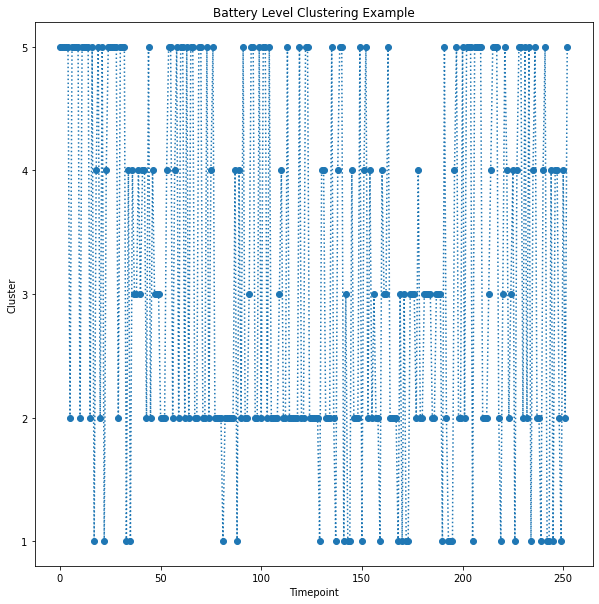
Timeseries Clustering¶
The example code:
"""
Created on Tue Apr 6 14:01:11 2021
@author: arsii
"""
import pandas as pd
import numpy as np
from tscfat.Analysis.cluster_timeseries import cluster_timeseries
# Load the data and convert index into Datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/Battery_test_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
df = df.resample('H').mean()
df = df.interpolate()
df = df.resample('D').apply(list)
df = df[1:-1]
data = np.stack(df.battery_level.values)
clusters = cluster_timeseries(data,
FIGNAME = False,
FIGPATH = False,
title = 'Battery Level',
n = 5,
mi = 5,
mib = 5,
rs = 0,
metric = 'dtw',
highlight = None,
ylim_ = (0,100)
)
The output images:

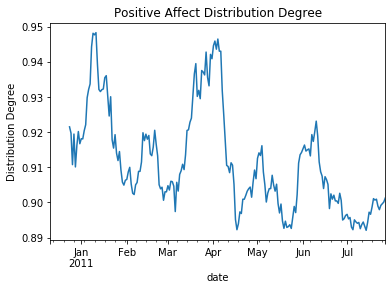
Degree of Distribution¶
The example code:
"""
Created on Mon Apr 5 21:18:34 2021
@author: arsi
"""
import pandas as pd
from tscfat.Analysis.degree_of_distribution import distribution_degree
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# set range of fluctuation
scale = 1
# window
w = 14
# calculate distribution degree
dist_deg = df.positive.rolling(window = w).apply(lambda x: distribution_degree(x.values,scale,w))
# plot the result
ax = dist_deg.plot()
ax.set(ylabel='Distribution Degree', title="Positive Affect Distribution Degree")
The output image:
Fluctuation Intensity¶
The example code:
"""
Created on Sat Apr 3 23:48:38 2021
@author: arsi
"""
import pandas as pd
from tscfat.Analysis.fluctuation_intensity import fluctuation_intensity
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# set range of fluctuation
scale = 1
# window
w = 14
# calculate fluctuation intensity
flu_int = df.positive.rolling(window = w).apply(lambda x: fluctuation_intensity(x.values,scale,w))
# plot the result
ax = flu_int.plot()
ax.set(ylabel='Fluctuation Intensity', title="Positive Affect Fluctuation Intensity")
The output image:

Plot Timeseries¶
The example code:
"""
Created on Fri Apr 2 12:14:27 2021
@author: arsi
"""
import pandas as pd
from tscfat.Analysis.plot_timeseries import plot_timeseries
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# A list containing column names
cols = ['positive','negative']
# Rolling window size
window = 14
_ = plot_timeseries(df,
cols,
title = 'Positive and negative affects',
roll = window,
xlab = "Time",
ylab = "Value",
ylim = False,
savename = False,
savepath = False,
highlight = False,
test = False
)
The output image:
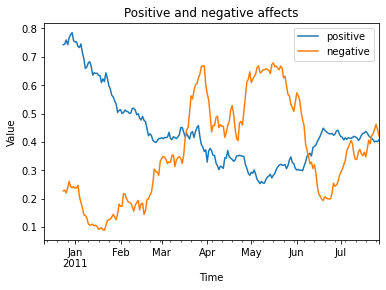
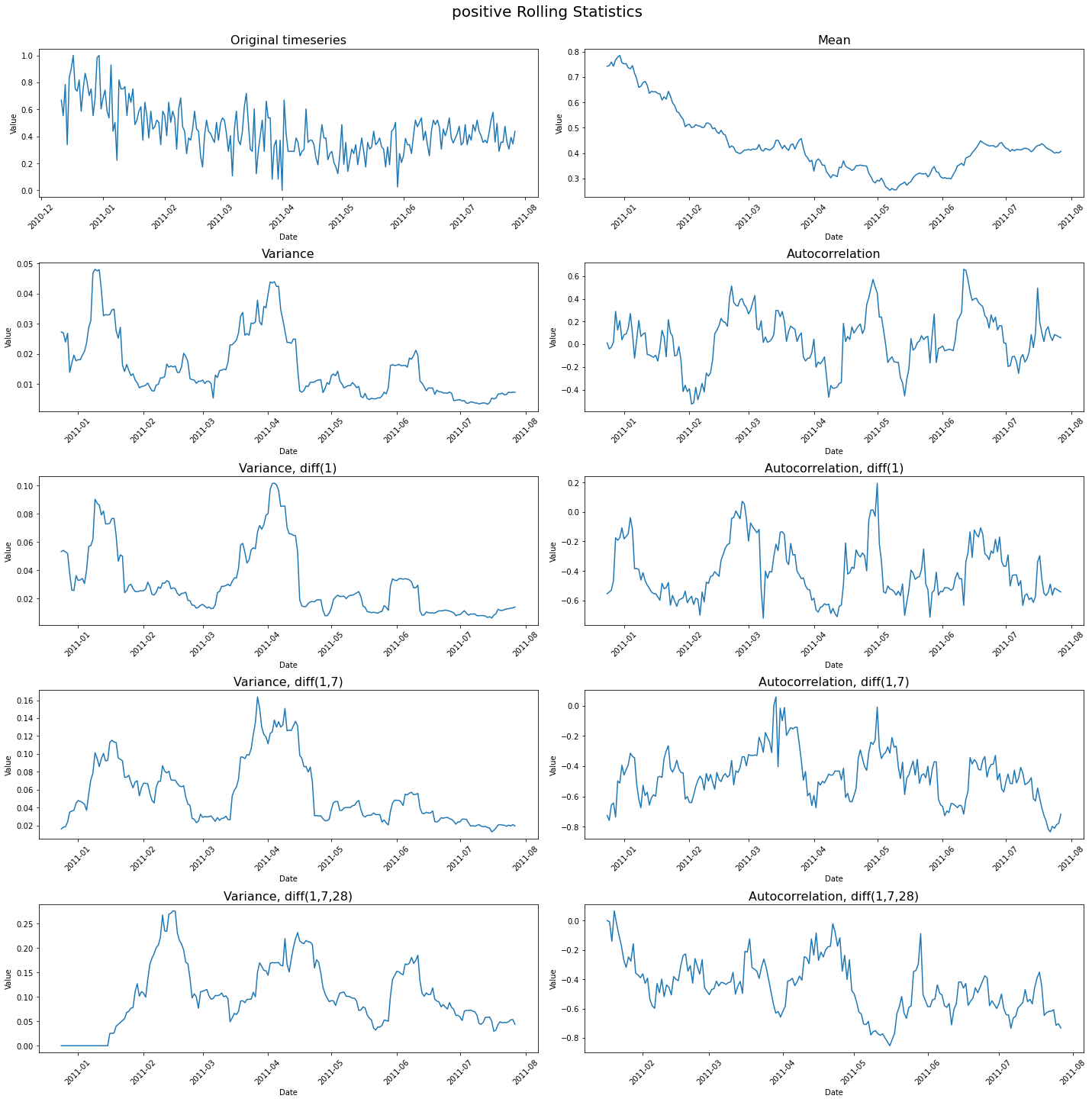
Rolling Statistics¶
The example code:
import pandas as pd
from tscfat.Analysis.rolling_statistics import rolling_statistics
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsii/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# dataframe can contain only one column
df = df.filter(['positive'])
# rolling window length
window = 14
_ = rolling_statistics(df,
window,
doi = None,
savename = False,
savepath = False,
test = False,
)
The output image:
Summary Statistics¶
The example code:
"""
Created on Mon Apr 5 21:40:44 2021
@author: arsi
"""
import pandas as pd
from tscfat.Analysis.summary_statistics import summary_statistics
# load a dataframe and convert the index to datetime
df = pd.read_csv('/home/arsi/Documents/tscfat/Data/one_subject_data.csv',index_col=0)
df.index = pd.to_datetime(df.index)
# Select 'Positive' column and convert it into pandas series
ser = df['positive']
# Rolling window size
w = 14
_ = summary_statistics(ser,
title = "Time series summary",
window = w,
savepath = False,
savename = False,
test = False,
)
The output image:
